Regina Zopf, FDA, Bethesda, MD, USA
Jeff Abolafia, Rho, Chapel Hill, NC, USA
Bhargava Reddy, UCB, Raleigh, NC USA
ABSTRACT
There is a lot of interest in the clinical trial community to understand what information can be obtained from Electronic Health Records (EHRs) to support clinical trials. The use of FHIR has been endorsed by the Office of National Coordinator for Health Information Technology (ONC) and is widely being used by EHR vendors. A pilot was conducted to assess whether data that are typically of interest (commonly collected) in the treatment of diabetic patients can be extracted from medical records through FHIR resources to support data collection for clinical trials. Specifically, we assessed data in the SyntheticMass’ Synthea repository that could be used to populate the demographics, diabetes diagnosis, medical history, concomitant medications, vital signs and a selection of laboratory results.
Deliverables from this pilot include a sample annotated Case Report Form (CRF) with Clinical Data Acquisition Standards Harmonization (CDASH), Study Data Tabulation Model (SDTM) and FHIR Resource Path annotations; sample datasets; a summary of our process followed and pilot experience, including challenges we encountered.
INTRODUCTION
The purpose of this pilot was to test the use of Health Level 7 International’s (HL7) FHIR standard using a synthetic EHR database to generate RWE consistent with data standards now required by the United States Food and Drug Administration (FDA) and Japan’s Pharmaceuticals and Medical Devices Agency (PMDA). The pilot also served to assess the level of harmonization between FHIR and Clinical Data Interchange Standards Consortium (CDISC) CDASH data collection and SDTM data reporting standards.
BACKGROUND
REAL WORLD EVIDENCE
RWE can come from a variety of sources including EHRs, payer administration claims, and patient registries (Sherman, 2016). One of the challenges in obtaining RWE from existing data sources stems from the diverse array of applications used to collect and store healthcare data. The HL7 FHIR standard can serve as an interface between diverse EHR applications.
HL7 FHIR
HL7 (a health data Standards Development Organization (SDO)) published FHIR Release 3 in March 2017. FHIR is a standard for exchanging healthcare information electronically that defines an Application Programming Interface (API) for exchanging data between healthcare applications, particularly EHRs. The basic building block in FHIR is a resource and all exchangeable content is defined as a resource. Resources all share the following set of characteristics:
- A common way to define and represent them, building them from data types that define common reusable patterns of elements
- A common set of metadata
- A human readable part
FHIR's interoperability across various data sources makes it a logical choice for facilitating the pre-population of clinical research eCRFs from EHR data. Pre-populating the eCRF using existing EHR data is also known as eSource (CDISC, 2006 & Mitchel, 2015). Researchers have tested a number of different approaches for eSource (Hume, et al. , 2016) including many that use the CDISC Operational Data Model (ODM) standard. FHIR provides an alternative and potentially more effective standard for accessing and integrating RWE into new drug or biologic licensing applications.
Overall, FHIR aims to address the common 80% of healthcare data exchange needs, while leaving the other 20% to extensions. (FHIR, 2017) Not all the information needed to complete every eCRF will be available in the basic FHIR resources, but extensions can be applied to improve the coverage.
FHIR REPRESENTATIONAL STATE TRANSFER (RESTful) API
HL7 developed a Representational State Transfer (RESTful) API for FHIR using existing internet technologies. RESTful APIs have broad acceptance at technology organizations like Google, Facebook, Twitter, Amazon, and numerous others. The API form of data interchange has been labeled the "API Economy" by some (Vukovic et al., 2016). FHIR brings the "API Economy" to healthcare by allowing healthcare data to be integrated from diverse organizations and technologies. Integration of healthcare data using the FHIR API offers the potential to deliver integrated healthcare solutions much faster than previously possible.
SYNTHEA DATABASE
Synthetic Mass is an open-source, simulated Health Information Exchange (HIE) populated with EHR data from one million realistic “synthetic residents” of Massachusetts. The synthetic population in the SyntheticMass “Synthea” database statistically mirrors the real population in terms of demographics, disease burden, vaccinations, medical visits, and social determinants. The database has implemented the FHIR API and thus the data can be accessed using the FHIR standard.
CDISC
CDISC is an SDO that has developed data standards used across the biopharmaceutical industry, enabling the harmonization of clinical trial data. CDISC provides guidelines for data collection (CDASH) as well as for reporting clinical trial data to regulatory authorities (SDTM and Analysis Data Model (ADaM)). The use of standards provides traceability from the planning phase, through data collection, tabulation and statistical analysis. The CDISC Coalition for Accelerating Standards and Therapies (CFAST) initiative aided in the creation of Therapeutic Area User Guides (TAUG). These TAUGs define data standards for core sets of clinical concepts and endpoints for targeted therapeutic areas. The Diabetes TAUG provides a core set of standards related to treating diabetes in adult outpatient populations. The Diabetes TAUG includes subject and disease characteristics, disease assessments and routine data.
METHODS
We defined selection criteria to identify adult subjects with Diabetes Mellitus Type 2 with at least two encounters in the Synthea Database. These are outlined in the table below.
| Selection Criteria |
|---|
| Age >18 years of Age |
| Diagnosis of Type 2 Diabetes Mellitus (ICD-10 Code E11.n) |
| Subject has at least two encounters in the Synthea database |
| CDASH Data Fields Extracted |
| Unique identifier |
| Characteristics: Age, Sex, Weight, BMI |
| Concomitant Medications |
| Encounter Type |
| Labs: Hemoglobin A1C, Fasting Plasma Glucose, Creatinine (Diabetes TAUG) |
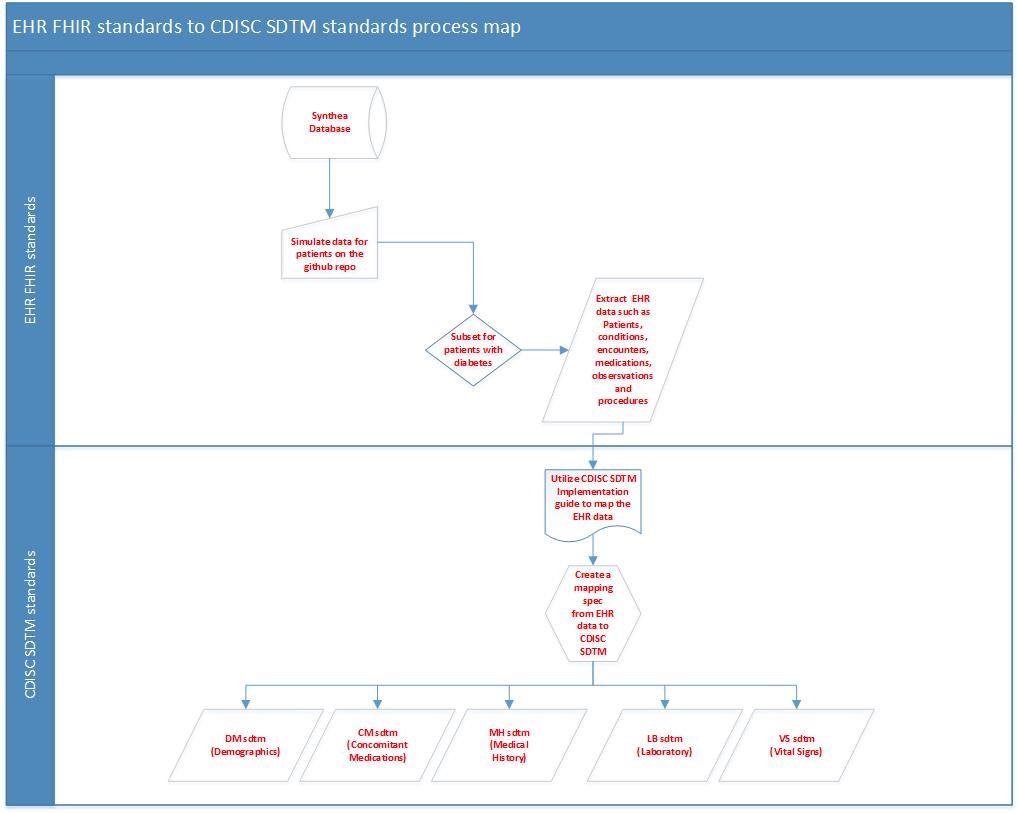
We accessed the SyntheticMass Synthea database (Synthea Generic Module Framework, 2017) and extracted Synthea data output in Java Script Object Notation (JSON) and Comma Separated Value (CSV) formats. Extracted data points included patient level medical records such as demographics, encounter information, medications, medical history, laboratory and vital signs, as shown in Figure 1.
Patient EHR data is generated using the Synthea database by simulation. The simulation is performed using Ruby script and cloning the Synthea database available on the GitHub repository. Ruby 2.1.0 or above is required to clone selected patient data from the Synthea database. Using Ruby programming, we then generated a dataset of synthetic patient data by filtering for the disease condition “Diabetes”. The FHIR standard data we obtained was in a CSV format and included data from patients of all ages with both Type 1 and Type 2 diabetes mellitus. Therefore, we used SAS code to further select the desired patients for inclusion in our final dataset. We then evaluated the data to ensure it was complete and ready to be mapped to CDISC SDTM variables. In the future, a FHIR API should allow users to select directly from FHIR data according to specific selection criteria.
MAPPING FROM THE FHIR STANDARD TO CDASH AND SDTM
A major objective of this project was to map data from the FHIR standard to the CDISC CDASH and SDTM standards (see Figure 1). Another objective was to assess the degree of harmonization between FHIR and the CDASH/STDM CDISC standards. The FHIR and CDISC standards organize data or information differently. In CDISC, information is grouped into a series of "data domains", such as demography or vital signs. In contrast, FHIR information is grouped into a collection of "resources", each containing a type of clinical or administrative information, such as prescriptions or procedures.
The data fields we selected for the pilot came from the Demography, Vital Signs, Laboratory, Concomitant Medications, and Medical History CDASH CRFs. We extracted all concomitant medications and medical history regardless of the time period for which they were available. All data fields extracted are listed in Table 1 above.
We created an SDTM mapping specification document to delineate the equivalent mapping variable for each data point available in the Synthea EHR FHIR dataset. We performed the mapping manually to match each FHIR resource path to CDISC SDTM data elements according to the relevance of the content. We used controlled terminology wherever required in order to be consistent with the CDISC Diabetes TAUG for CDISC SDTM data mapping. Table 2 displays the data elements we selected and the respective variables in the FHIR resource path, CDASH, and SDTM.
Demography was the first data domain we evaluated. The demography variables of interest in FHIR were all located in the FHIR Patient resource. The Patient resource contains demographic and other administrative information about a patient. In SDTM/CDASH the unique subject identifier is stored in the single field, USUBJID, while in FHIR a unique patient is identified by a patient ID (Identifier.value) and a system identifier (Identifier.system). De-identification of data was unnecessary for the synthetic data obtained for the purposes of this pilot. We acknowledge that in the future when real EHR data are used to populate datasets, a de-identifying process will be needed.
Note: In FHIR, multiple data elements are formed into resources that are arranged in a hierarchical manner and therefore, to refer to individual elements, a convention of element.subelement is used with the Resource name being referenced in the text. The subelement part can repeat. Full path names are provided in the FHIR Resource Path column in Table 2.
For the SDTM variable SEX, the FHIR variable gender was mapped to the SDTM standard. We recognize gender and sex may not be the same in all cases. Date/time of birth was similar in FHIR and SDTM, but required reformatting for CDASH. In general, date/times in FHIR are stored in the ISO 8601 format. When converting a FHIR data to CDASH, it is split into separate data and time components. Dates in SDTM are then constructed from the separate CDASH data and time components. In FHIR, AGE was not a defined concept but could easily be derived at a given time point from the Date/time of birth resource. In CDASH/SDTM, AGE is either collected or derived.
The FHIR Condition resource contains information about a condition, problem, or diagnosis and therefore was used to identify subjects with Diabetes Mellitus Type 2. In CDASH/SDTM this information is stored in the Medical History domain. In our pilot database, the FHIR variables coding.code and coding.display contain the SNOMED code and associated text description for a given condition. These two variables map to the CDASH/SDTM fields MHDECOD and MHTERM, respectively. It is important to note that the Condition resource contains a field called coding.system that links to the coding dictionary used. As a result, FHIR is not limited to a single coding system. FHIR specifies fields for date of assessment (assertedDate), start date of condition (onsetDateTime), and end date of condition (abatementDateTime) that can be mapped directly to its SDTM counterparts MHDTC, MHSTDTC, MHENDTC and can be reformatted to CDASH dates as described above. Category of medical history is stored in the FHIR field category, which corresponds to the CDASH/SDTM MHCAT variable. The CDASH/SDTM concepts Medical History Event Pre-Specified (MHPRESP) and Medical History Occurrence (MHOCCUR) are not represented (or applicable) in FHIR.
A third area of interest was concomitant medications. In FHIR, medication information is contained in the Medication and Medication Statement resources. While the Medication resource is used to identify and define a medication, the Medication Statement resource provides information about medications that are administered to a patient. The Medication resource contains fields for medication code (coding.code) and the name of the medication (coding.display). These two fields correspond to the CDASH/SDTM fields CMDECOD and CMTRT. We used the Synthea data verbatim; we did not recode. As in the Condition resource, the Medication resource also contains a field containing a link to the coding dictionary used and is therefore not limited to a single coding dictionary. In FHIR, additional details about the medication are located in the Medication Statement resource. The amount of dose and dose unit are stored in the FHIR fields doseage.doseQuantity.value and doseage.doseQuantity.code. These two fields can be mapped to the CDASH/SDTM variables CMDOSE and CMDOSU. Dosing frequency and route of administration are housed in the FHIR variables Doseage.timing and Doseage.route, which correspond to the CDASH/SDTM variables CMDOSFRQ and CMROUTE. In FHIR, start and end dates for medication are represented by effectivePeriod.start and effectivePeriod.end. These two fields map directly to the SDTM variables CMSTDTC and CMENDTC and can be reformatted to the CDASH fields CMSTDAT and CMEDDAT. The CDASH concept of “Ongoing” does not have a counterpart in FHIR. However, “Ongoing” can be derived: if start date is present and end date is missing, then the medication is considered to be ongoing for a given encounter.
Height, Weight, BMI, and Laboratory test results are all represented in the Observation resource, which contains measurements and assertions made about a patient, and in the CDASH/SDTM Vital Signs (VS) and Laboratory Test Results (LB) domains. Since LB and VS are both in the general observation class “Findings”, they are mapped to FHIR in a similar manner. Laboratory and vital sign parameters are specified in the Synthea FHIR data by a LOINC code (coding.code) and an associated text description (coding.display). A given LOINC code provides a description of the parameter, the unit of measurement, the specimen type, the method of the test and the category for the test or vital sign. We used the unit field Observation.valueQuantity. However, one could use the unit found in the specific LOINC code. A FHIR LOINC code and description can be mapped to the STDM/CDASH fields test name (LBTEST/VSTEST) and short test name (LBTESTCD/VSTESTCD). Controlled terminology is likely to differ between the LOINC code descriptions and CDISC controlled terminology (See Table 3).
For each laboratory and vital signs parameter, Error! Reference source not found. shows the associated LOINC code, short and long descriptions, and corresponding CDISC controlled terminology for LBTEST/VSTEST and LBTESTCD/VSTESTCD. In most cases LOINC code descriptions are not harmonized with CDISC controlled terminology. LOINC codes can be stored in the CDASH/SDTM fields LBLOINC/VSLOINC. The result or finding in FHIR is stored in the variable valueQuantity.value, while its CDASH/SDTM counterpart is housed in LBORRES/VSORRES. The unit of measurement in CDASH/SDTM (LBORRESU/VSORRESU) can be ascertained by its LOINC code. Additionally, for lab parameters, the specimen type (LBSPEC), the method of test (LBMETHOD), and category for lab test (LBCAT) can be ascertained from the LOINC code. Lastly, date of examination is represented in FHIR as effectiveDateTime and LBDAT/VSDATA in CDASH and LBDTC/VSDTC in SDTM.
The final concept of interest was encounter type. The FHIR Encounter resource contains extensive information about the patient/health care provider interaction. The encounter type is stored the field “class” and documents where the encounter took place (i.e. emergency, inpatient, home health, etc.). We used the encounter type as part of patient selection criteria for this pilot. CDASH/SDTM does not currently have a specific variable encounter type. This could present a challenge to researchers using RWE if the information is considered relevant for a specific research question.
| Domain | Concept | FHIR RESOURCE PATH (Resource.Element.Subelement) | CDASH Variable | SDTM Variable | Notes |
|---|---|---|---|---|---|
| DM | Unique Subject ID | Patient.Identifier.value Patient.Identifier.system | USUBJID | USUBJID | In FHIR a unique patient is identified by a patient ID and a system identifier |
| Sex | Patient.Gender | SEX | SEX | When converting to CDISC, gender must be re-coded | |
| Age | n/a | AGE | AGE | In FHIR, age is calculated for birth date and a reference date | |
| Birth Date | Patient.birthDATE | BRTHDAT | BRTHDTC | Date/times in FHIR are stored in ISO 8601. Date/times in STDM are also stored in ISO 8601 format, but typically constructed from CDASH date/time components | |
| MH | Verbatim or pre-printed term | Condition.coding.display | MHTERM | MHTERM | |
| Coded Term | Condition.coding.code | MHDECOD | |||
| MHDECOD | |||||
| Category | Condition.category | MHCAT | MHCAT | ||
| Start Date | Condition. | ||||
| onsetDateTime | |||||
| MHSTDAT | MHSTDTC | ||||
| End Date | Condition. | ||||
| abatementDateTime | |||||
| MHENDAT | MHENDTC | ||||
| Collection Date | Condition.assertedDate | ||||
| MHDAT | MHDTC | ||||
| Pre-specified | n/a | MHPRESP | |||
| MHPRESP | |||||
| Occurrence | n/a | MHOCCUR | |||
| MHOCCUR | |||||
| CM | Name of Medication | Medication.display | CMTRT | CMTRT | |
| Standardized Name | Medication.display | CMDECOD | CMDECOD | ||
| Amount of Dose | Medication Statement. | ||||
| doseage.doseQuantity.value | |||||
| CMDOSE | CMDOSE | ||||
| Dose Unit | Medication Statement. | ||||
| doseage.doseQuantity.value | CMDOSU | CMDOSU | |||
| Frequency | Medication Statement. | ||||
| doseage.timing | CMDOSFRQ | ||||
| CMDOSFRQ | |||||
| Route | Medication Statement. | ||||
| doseage.route | CMROUTE | CMROUTE | |||
| Start Date | Medication Statement. | ||||
| effectivePeriod.start | |||||
| CMSTDAT | CMSTDTC | ||||
| End Date | Medication Statement. | ||||
| effectivePeriod.start | CMENDAT | CMENDTC | |||
| Ongoing | n/a | CMONGO | CMENRF | CMENRF derived from CMONGO | |
| VS/LB | |||||
| Test Name | Observation.coding.display | xxTEST | xxTEST | xx refers to LB or VS | |
| Short Name | n/a | xxTESTCD | xxTESTCD | ||
| Category | n/a | xxCAT | xxCAT | Derived from LOINC | |
| Result or Finding | Observation.valueQuantity. | ||||
| value | |||||
| xxORRES | xxORRES | ||||
| Unit | Observation.valueQuantity. | ||||
| Unit | |||||
| xxORRESU | xxORRESU | ||||
| LOINC Code | Observation.coding.code | xxLOINC | xxLOINC | ||
| Specimen Type | Derived from LOINC code | LBSPEC | LBSPEC | LBSPEC derived from LBLOINC | |
| Method Test | Derived from LOINC code | LBMETHOD | LBMETHOD | LBSPEC derived from LBLOINC | |
| Collection Date | Observation. | ||||
| effectiveDateTime | xxDAT | xxDTC | |||
| Encounter | Encounter Type | Encounter.class | n/a | n/a | Not a CDASH/SDTM concept |
| LOINC Code | LOINC Description | LOINC Short Description | LBTEST/VSTEST | LBTESTCD/VSTESTCD |
|---|---|---|---|---|
| 2339-0 | Glucose [Mass/volume] in | |||
| Blood | ||||
| Glucose Bld-mCnc | Glucose | GLUC | ||
| 4548-4 | Hemoglobin A1c/Hemoglobin.total in Blood | Hgb A1c MFr Bld | Hemoglobin A1C/Hemoglobin | |
| HBA1CHGB | ||||
| 14959-1 | Microalbumin/Creatinine [Mass Ratio] in Urine | Microalbumin/Creat Ur | Albumin/Creatinine | ALBCREAT |
| 33914-3 | Glomerular filtration rate/1.73 sq M.predicted [Volume Rate/Area] in Serum or Plasma by Creatinine-based formula (MDRD) | |||
| GFR/BSA.pred SerPl MDRD-ArVRat | Glomerular Filtration Rate, Estimated | GFRE | ||
| 8302-2 | Body Height | Height | Height | Height |
| 29463-7 | Body Weight | Weight | Weight | Weight |
| 39156-5 | Body mass index (BMI) Ratio | BMI | Body Mass Index | BMI |
TESTING THE FHIR RESTful API
To test the retrieval of Synthea data using the FHIR API, a Healthcare Services Platform Consortium (HSPC) sandbox (Vukovic et al., 2016) was instantiated to create the FHIR server needed to retrieve resources as well as to pre-populate the sandbox with Synthea data. After instantiating the sandbox, tests were developed and executed to evaluate the FHIR API’s ability to identify patients that matched the previously stated inclusion criteria and to retrieve the JSON-formatted FHIR resources previously listed for use in this study. Test scripts were developed in Python using the SMART on FHIR-based fhirclient ( HSPC, 2017). These tests evaluated the feasibility of using the FHIR API to retrieve EHR data for subsequent loading into CDISC CDASH eCRFs.
SUMMARY OF RESULTS/FINDINGS
MAPPING FROM THE FHIR STANDARD TO CDASH AND SDTM
We successfully mapped all variables of interest from FHIR resource paths to CDASH/SDTM eCRF pages/fields of interest for this pilot with the exception of encounter type. While age needs to be calculated in FHIR, it is often derived in CDASH/SDTM as well. Encounter type is a FHIR concept, but does not have a CDASH/SDTM counterpart. Although most of the variables of interest are represented in both FHIR resource paths and CDASH/SDTM, in many cases they lack harmonization. All concepts have variable names that are different in FHIR and CDASH/SDTM. Furthermore, many of our variables have controlled terminology that is not harmonized. Further work is needed by the CDISC EHR-to-CDASH (E2C) team to develop standard mappings.
It is important to note that medications and medical diagnoses were only available in SNOMED terminology and ICD-10 coding in the SyntheticMass database. This is consistent with most EHR systems that commonly capture diagnoses using ICD-10 coding for billing purposes. In the pharmaceutical industry, adverse event data related to drug exposure is provided to regulatory authorities using MedDRA terminology and for medications using the WHO Drug Dictionary. A new variable to capture the original SNOMED or ICD-10 terminology would need to be created along with an explanation of the process to convert SNOMED/ICD-10 to MedDRA and/or WHO Drug coding. A standard approach to “translation” between dictionaries would greatly enhance the ability to use RWE in drug development.
TESTING THE FHIR RESTful API
The Python test scripts demonstrated that FHIR Resource data for diabetes patients in an EHR could be identified and retrieved using the FHIR API with software tools based on existing libraries.
Snapshots of the final SDTM output and an example eCRF with annotated data elements are provided in the Appendix.
DISCUSSION
This pilot project was an important first step in the exploration of the use of FHIR in the generation of CDISC compliant real world evidence. Further work is needed in order to fully realize the potential benefits that FHIR offers in the use of RWE but the concept shows great promise to enable the collection and use of real world data. While it is possible to map EHR data using the FHIR standard to CDASH compliant outputs, there are limitations to the use of this data in a real world clinical trial setting. Examples of these limitations include missing and/or incomplete data, irregular and inconsistent data collection across hospitals/clinical sites and heterogeneity in dictionaries used in the original data collection. The applicability of our project to real world data is limited by the narrow scope and the small sample of data domains and variables. The Synthea database may not reflect “typical” EHR source data. FHIR does not provide metadata or question text for a given concept as compared to CDASH. This was not a concern for the variables we chose to explore for this project but it may be a concern for other variables.
The RESTful API developed as part of the FHIR specification provides an implementation friendly mechanism for retrieving FHIR Resources that distinguishes it from many previous healthcare standards. Much like many other data-centric businesses that have developed thriving RESTful APIs for data exchange, the availability of the FHIR API lowers the implementation burden for retrieving EHR data for secondary use.
The FHIR API offers a way forward building on the work performed for this pilot. For the pilot the data for a large number of patients was obtained in bulk, the selection criteria applied and the data for the matching patients/subjects extracted to buildthe SDTM domains.
In the future, we can envisage using FHIR to actually query the exact data that we need. We could query the EHR to provide a list of subjects matching the criteria and then, using the CRF as the specification, query those matching subjects forobservations matching the required data. For example, a vital signs form may only require Height, Weight and Blood Pressure and therefore we only query for those specific observations. We can be more precise in our data requests.Taking the vision further, queries like those described could be performed in real-time if such access was considered beneficial. We can also envisage the definition of a set of 1 or more FHIR queries that could provide the raw data for a givenSDTM domain such as demographics.
CONCLUSION
Electronic Case Report Form pages/fields of interest can be populated using EHR data by mapping FHIR concepts toCDASH/SDTM variables from the CDISC Diabetic TAUG.
To make FHIR to CDISC mapping more useful, it is recommended that we develop code that performs real-time translation. The PhUSE team hopes to continue our work by creating template working code that ingests data in FHIR format, and outputsdata in CDISC/ODM/CDASH and SDTM formats for consumption by clinical systems. The objective is to use open exchanges such as GitHub, to publish mapping documents and template translation code that can leverage standard API’s from EMR systems, EDC systems and other clinical systems.
REFERENCES
Interchange, C. E. S. D. Group. Leveraging the CDISC standards to facilitate the use of electronic source data within clinical trials version, November 20, 2006.
Healthcare Services Platform Consortium Sandbox Manager. Retrieved September 01, 2017, from https://sandbox.hspconsortium.org/
Hume, S., Aerts, J., Sarnikar, S., & Huser, V. (2016). Current applications and future directions for the CDISC OperationalData Model standard: A methodological review. Journal of biomedical informatics, 60, 352-362.
Mitchel, J. T., Helfgott, J., Haag, T., Cappi, S., McCanless, I., Kim, Y.J., Choi, J., Cho, T., Gittleman, D.A. (2015). eSource Records in Clinical Research. Applied Clinical Trials.
Synthea Generic Module Framework., Retrieved August 01, 2017, from https://github.com/synthetichealth/synthea/wiki/Generic-Module-Framework.
Vukovic, M., Laredo, J., Muthusamy, V., Slominski, A., Vaculin, R., Tan, W., Naik, V., Silva-Lepe,I., Srivastava,B., & Branch, J. W. (2016). Riding and thriving on the api hype cycle. Communications of the ACM, 59(3), 35-37. FHIR: Fast healthcare interoperability resources.Retrieved August 01, 2017 from http://hl7.org/implement/standards/fhir/
APPENDIX
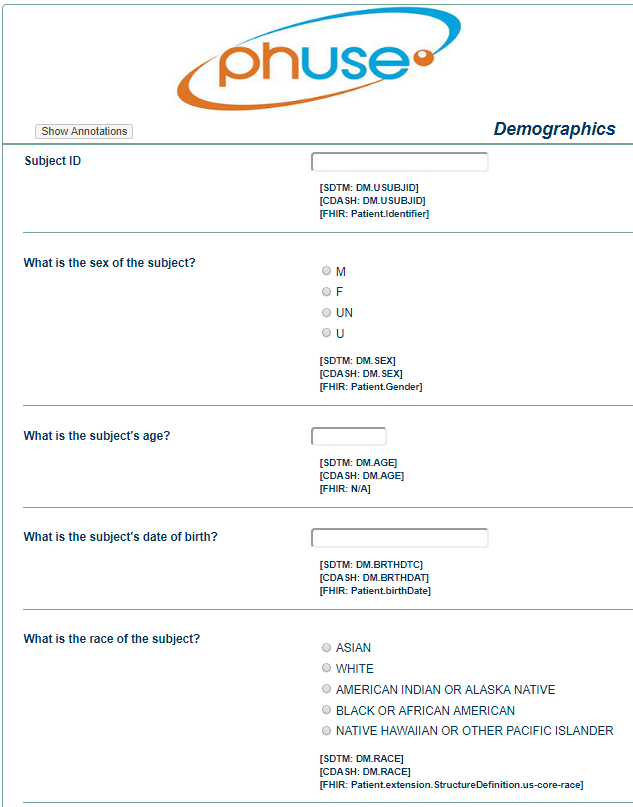
DEMOGRAPHY CRF WITH ANNOTATIONS
